AI Just Crushed a Major Barrier in Biomedical Research!
Authors:
Youngjun Park (FAIrPaCT Member)
Nils P. Muttray
Anne-Christin Hauschild (FAIrPaCT Member)
Published in:
Briefings in Bioinformatics, 2024
👉 Read the paper here : https://academic.oup.com/bib/article/25/2/bbae004/7596256?login=true
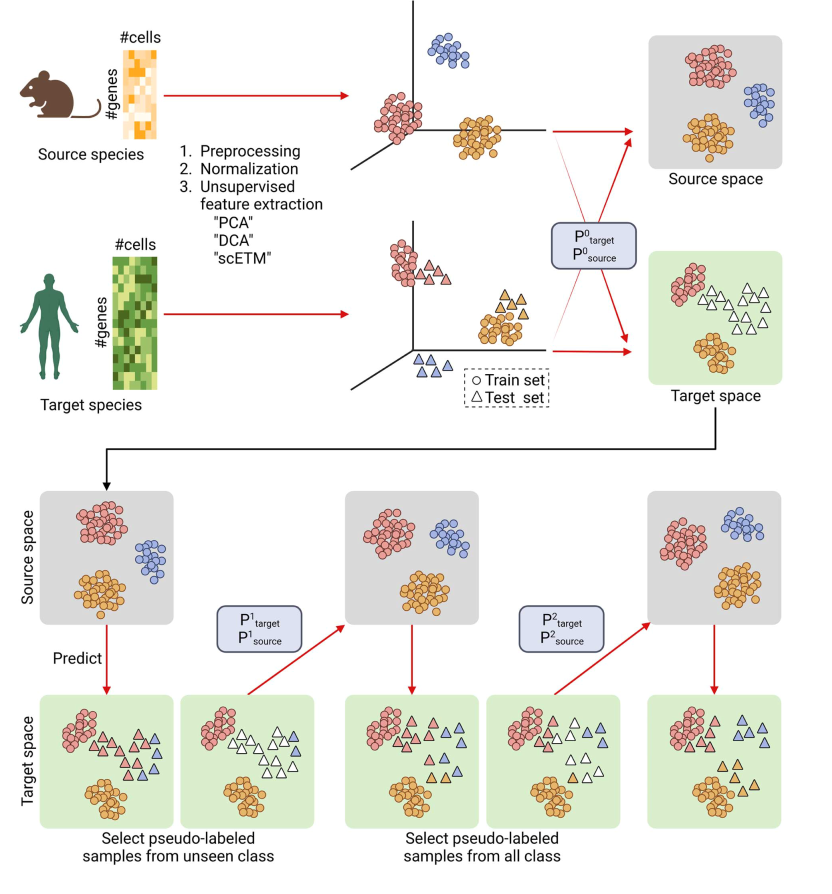
different domains into a common latent space with a projective matrix (Psource/target). The latent space is further aligned with two pseudo-labeling steps. During the first pseudo-labeling step, the latent space is updated with the train set and only unseen classes of the test set. The second pseudo-labeling step utilizes all classes in the test set for the pseudo-labeling and further updates the alignment of latent spaces. In this common space, unlabeled data in the target species can be predicted with knowledge of the source species.
What’s the Big Deal?
Biomedical research relies heavily on model organisms (think: mice, zebrafish) to study human diseases. The problem? Different species = different genes. Translating discoveries from animals to humans has always meant navigating messy gene orthology databases—losing valuable information in the process.
But FAIrPaCT just helped rewrite the rules.
Meet SATL: Species-Agnostic Transfer Learning
💡 No more gene-ID conversion. No more info loss.
Our team developed SATL, a cutting-edge machine learning approach that connects datasets from entirely different species—WITHOUT needing gene orthology. SATL learns a shared biological “language” across species and predicts unseen cell types with surprising accuracy.
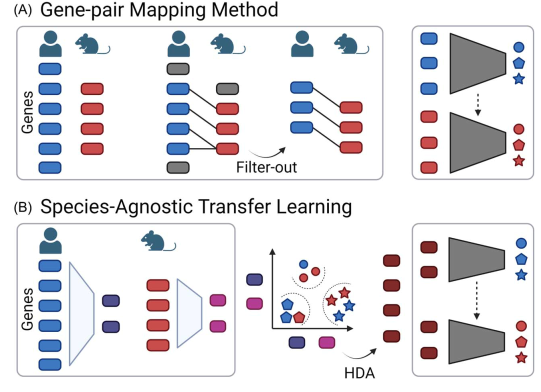
extractor is used to embed a gene set into latent features. After feature extraction, heterogeneous features are aligned using an HDA algorithm. The aligned features can then be used directly in the integrative analysis.
🔥 Why This Rocks
✅ Cross-species predictions that actually work – SATL crushes it on human-mouse single-cell datasets from bone marrow, pancreas, and brain.
✅ No external databases required – say goodbye to tedious gene mapping.
✅ Preserves biological meaning – finds immune system and other key biological pathways shared across species.
✅ Beats standard tools – outperforms methods like Mutual Nearest Neighbors (MNNs) and CADA-VAE, even in zero-shot scenarios.
Why It’s Pure FAIrPaCT Energy
This is AI for good. SATL helps researchers unlock hidden knowledge across species without compromising data integrity. It’s privacy-conscious, efficient, and totally aligned with our mission to power ethical and high-impact AI in life sciences.
🌍 From lab bench to bedside, SATL brings us one step closer to smarter, faster biomedical breakthroughs.
Stay tuned for more FAIrPaCT-driven innovations! 🚀✨
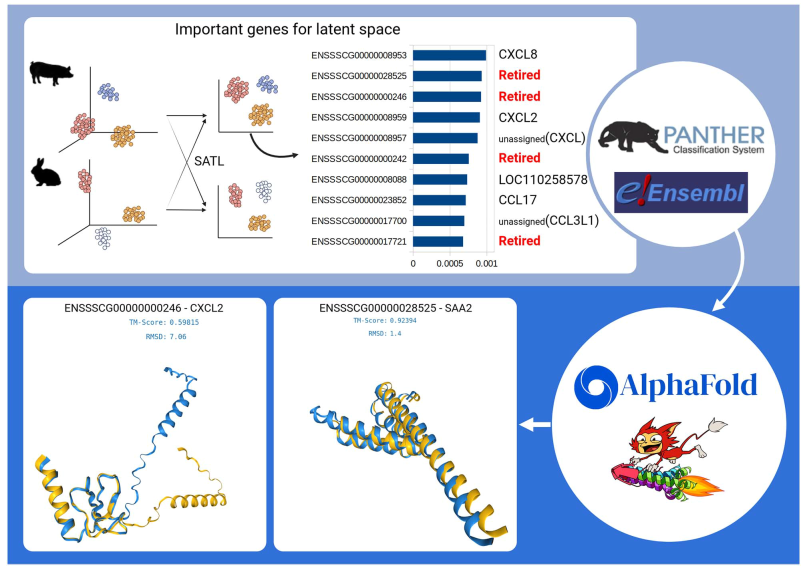
current orthologous database. Two genes are retired in the Ensembl database. Therefore, structural similarity analysis of protein is done. Foldseek search identified ‘ENSSSCG00000028525’ as a similar protein to CXCL2 with sequence identity: 75.7 and E-Value: 3.87e−14. ‘ENSSSCG00000000246’ is identified as a similar protein to SAA2 with sequence identity: 72.3 and E-Value: 2.47e-13. ‘ENSSSCG00000017721’ and ‘ENSSSCG00000000242’ are identified as ‘CCL8’ and ‘CCL2’.

